Abstract
Background
Lung adenocarcinoma (LUAD) is the most common histological type of lung cancer with lower survival rates. Recent advancements in targeted therapies and immunotherapies targeting immune checkpoints have achieved remarkable success, there is still a large percentage of LUAD that lacks available therapeutic options. Due to tumor heterogeneity, the diagnosis and treatment of LUAD are challenging. Exploring the biology of LUAD and identifying new biomarker and therapeutic targets options are essential.
Method
We performed single-cell RNA sequencing (scRNA-seq) of 6 paired primary and adjacent LUAD tissues, and integrative omics analysis of the scRNA-seq, bulk RNA-seq and whole-exome sequencing data revealed molecular subtype characteristics. Our experimental results confirm that CDC25C gene can serve as a potential marker for poor prognosis in LUAD.
Results
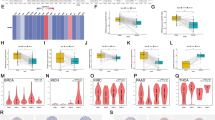
We investigated aberrant gene expression in diverse cell types in LUAD via the scRNA-seq data. Moreover, multi-omics clustering revealed four subgroups defined by transcriptional profile and molecular subtype 4 (MS4) with poor survival probability, and immune cell infiltration signatures revealed that MS4 tended to be the immunosuppressive subtype. Our study revealed that the CDC25C gene can be a distinct prognostic biomarker that indicates immune infiltration levels and response to immunotherapy in LUAD patients. Our experimental results concluded that CDC25C expression affects lung cancer cell invasion and migration, might play a key role in regulating Epithelial-Mesenchymal Transition (EMT) pathways.
Conclusions
Our multi-omics result revealed a comprehensive set of molecular attributes associated with prognosis-related genes in LUAD at the cellular and tissue level. Identification of a subtype of immunosuppressive TME and prognostic signature for LUAD. We identified the cell cycle regulation gene CDC25C affects lung cancer cell invasion and migration, which can be used as a potential biomarker for LUAD.







Similar content being viewed by others
Data availability
The experimental data that support the findings of this study are available in Figshare with the identifier https://doi.org/10.6084/m9.figshare.25027181.v1.
Abbreviations
- NSCLC:
-
Non-small cell lung cancer
- LUAD:
-
Lung Adenocarcinoma
- GEO:
-
Gene Expression Omnibus Database
- TCGA:
-
The Cancer Genome Atlas
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- ScRNA-seq:
-
Single-cell RNA sequencing
- TME:
-
Tumor microenvironment
- CNV:
-
Copy number variations
- EMT:
-
Epithelial-mesenchymal transition
- PDX:
-
patient-derived xenografts
References
The global challenge of cancer. Nat. Cancer 1, 1–2 (2020)
A. Leiter, R.R. Veluswamy, J.P. Wisnivesky, The global burden of lung cancer: current status and future trends. Nat. Rev. Clin. Oncol. 20, 624–639 (2023)
H. Sung, J. Ferlay, R.L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal, F. Bray, Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021)
J. Chen, H. Yang, A.S.M. Teo, L.B. Amer, F.G. Sherbaf, C.Q. Tan, J.J.S. Alvarez, B. Lu, J.Q. Lim, A. Takano, et al, Genomic landscape of lung adenocarcinoma in East Asians. Nat. Genet. 52, 177–186 (2020)
J.Y. Xu, C. Zhang, X. Wang, L. Zhai, Y. Ma, Y. Mao, K. Qian, C. Sun, Z. Liu, S. Jiang, et al, Integrative proteomic characterization of human lung adenocarcinoma. Cell 182, 245–261 e217 (2020)
M. Miyake, S. Hori, Y. Morizawa, Y. Tatsumi, Y. Nakai, S. Anai, K. Torimoto, K. Aoki, N. Tanaka, K. Shimada, et al, CXCL1-mediated interaction of cancer cells with tumor-associated macrophages and cancer-associated fibroblasts promotes tumor progression in human bladder cancer. Neoplasia 18, 636–646 (2016)
Y. Liang, Y. Tan, B. Guan, B. Guo, M. Xia, J. Li, Y. Shi, Z. Yu, Q. Zhang, D. Liu, et al, Single-cell atlases link macrophages and CD8(+) T-cell subpopulations to disease progression and immunotherapy response in urothelial carcinoma. Theranostics 12, 7745–7759 (2022)
L. Gonzalez-Silva, L. Quevedo, I. Varela, Tumor functional heterogeneity unraveled by scRNA-seq technologies. Trends Cancer 6, 13–19 (2020)
M.L. Metzker, Sequencing technologies - the next generation. Nat. Rev. Genet. 11, 31–46 (2010)
D. Sun, X. Guan, A.E. Moran, L.Y. Wu, D.Z. Qian, P. Schedin, M.S. Dai, A.V. Danilov, J.J. Alumkal, A.C. Adey, et al, Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data. Nat. Biotechnol. 40, 527–538 (2022)
A.S. Thind, I. Monga, P.K. Thakur, P. Kumari, K. Dindhoria, M. Krzak, M. Ranson, B. Ashford, Demystifying emerging bulk RNA-Seq applications: the application and utility of bioinformatic methodology. Brief Bioinform. 22, bbab259 (2021)
B. Jew, M. Alvarez, E. Rahmani, Z. Miao, A. Ko, K.M. Garske, J.H. Sul, K.H. Pietilainen, P. Pajukanta, E. Halperin, Accurate estimation of cell composition in bulk expression through robust integration of single-cell information. Nat. Commun. 11, 1971 (2020)
J. Liao, J. Qian, Y. Fang, Z. Chen, X. Zhuang, N. Zhang, X. Shao, Y. Hu, P. Yang, J. Cheng, et al, De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution. Nat. Commun. 13, 6498 (2022)
X. Guo, Y. Zhang, L. Zheng, C. Zheng, J. Song, Q. Zhang, B. Kang, Z. Liu, L. Jin, R. Xing, et al, Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 24, 978–985 (2018)
D. Lambrechts, E. Wauters, B. Boeckx, S. Aibar, D. Nittner, O. Burton, A. Bassez, H. Decaluwe, A. Pircher, K. Van den Eynde, et al, Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018)
J.D. Klement, P.S. Redd, C. Lu, A.D. Merting, D.B. Poschel, D. Yang, N.M. Savage, G. Zhou, D.H. Munn, P.G. Fallon, K. Liu, Tumor PD-L1 engages myeloid PD-1 to suppress type I interferon to impair cytotoxic T lymphocyte recruitment. Cancer Cell 41, 620–636 e629 (2023)
L. Zhang, Y. Zhang, C. Wang, Y. Yang, Y. Ni, Z. Wang, T. Song, M. Yao, Z. Liu, N. Chao, et al, Integrated single-cell RNA sequencing analysis reveals distinct cellular and transcriptional modules associated with survival in lung cancer. Signal Transduct. Target. Ther. 7, 9 (2022)
F. Wu, J. Fan, Y. He, A. Xiong, J. Yu, Y. Li, Y. Zhang, W. Zhao, F. Zhou, W. Li, et al, Single-cell profiling of tumor heterogeneity and the microenvironment in advanced non-small cell lung cancer. Nat. Commun. 12, 2540 (2021)
D. He, D. Wang, P. Lu, N. Yang, Z. Xue, X. Zhu, P. Zhang, G. Fan, Single-cell RNA sequencing reveals heterogeneous tumor and immune cell populations in early-stage lung adenocarcinomas harboring EGFR mutations. Oncogene 40, 355–368 (2021)
B. Liu, X. Hu, K. Feng, R. Gao, Z. Xue, S. Zhang, Y. Zhang, E. Corse, Y. Hu, W. Han, Z. Zhang, Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022)
H. Okayama, T. Kohno, Y. Ishii, Y. Shimada, K. Shiraishi, R. Iwakawa, K. Furuta, K. Tsuta, T. Shibata, S. Yamamoto, et al, Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res. 72, 100–111 (2012)
Y. Hao, T. Stuart, M.H. Kowalski, S. Choudhary, P. Hoffman, A. Hartman, A. Srivastava, G. Molla, S. Madad, C. Fernandez-Granda, R. Satija, Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. (2023)
Y. Hao, S. Hao, E. Andersen-Nissen, W.M. Mauck 3rd, S. Zheng, A. Butler, M.J. Lee, A.J. Wilk, C. Darby, M. Zager, et al, Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 e3529 (2021)
L.A. Huuki-Myers, K.D. Montgomery, S.H. Kwon, S.C. Page, S.C. Hicks, K.R. Maynard, Collado-Torres L: data-driven identification of total RNA expression genes for estimation of RNA abundance in heterogeneous cell types highlighted in brain tissue. Genome Biol. 24, 233 (2023)
S. Hanzelmann, R. Castelo, J. Guinney, GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013)
C.B. Thompson, K.H. Vousden, R.S. Johnson, W.H. Koppenol, H. Sies, Z. Lu, L.W.S. Finley, C. Frezza, J. Kim, Z. Hu, C.R. Bartman, A century of the Warburg effect. Nat. Metab. 5, 1840–1843 (2023)
P. Icard, S. Shulman, D. Farhat, J.M. Steyaert, M. Alifano, H. Lincet, How the Warburg effect supports aggressiveness and drug resistance of cancer cells? Drug Resist. Updat. 38, 1–11 (2018)
M. Fedele, R. Sgarra, S. Battista, L. Cerchia, G. Manfioletti, The epithelial-mesenchymal transition at the crossroads between metabolism and tumor progression. Int. J. Mol. Sci. 23, 800 (2022)
P. Sarode, S. Mansouri, A. Karger, M.B. Schaefer, F. Grimminger, W. Seeger, R. Savai, Epithelial cell plasticity defines heterogeneity in lung cancer. Cell. Signal. 65, 109463 (2020)
W.K. Cheung, D.X. Nguyen, Lineage factors and differentiation states in lung cancer progression. Oncogene 34, 5771–5780 (2015)
A.P. Patel, I. Tirosh, J.J. Trombetta, A.K. Shalek, S.M. Gillespie, H. Wakimoto, D.P. Cahill, B.V. Nahed, W.T. Curry, R.L. Martuza, et al, Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014)
I. Tirosh, B. Izar, S.M. Prakadan, M.H. Wadsworth 2nd, D. Treacy, J.J. Trombetta, A. Rotem, C. Rodman, C. Lian, G. Murphy, et al, Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016)
C. Trapnell, D. Cacchiarelli, J. Grimsby, P. Pokharel, S. Li, M. Morse, N.J. Lennon, K.J. Livak, T.S. Mikkelsen, J.L. Rinn, The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014)
M.D. Wilkerson, D.N. Hayes, ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010)
A.M. Newman, C.L. Liu, M.R. Green, A.J. Gentles, W. Feng, Y. Xu, C.D. Hoang, M. Diehn, A.A. Alizadeh, Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015)
J. Al-Matouq, T.R. Holmes, L.A. Hansen, CDC25B and CDC25C overexpression in nonmelanoma skin cancer suppresses cell death. Mol. Carcinog 58, 1691–1700 (2019)
N. Xu, Y. Ren, Y. Bao, X. Shen, J. Kang, N. Wang, Z. Wang, X. Han, Z. Li, J. Zuo, et al, PUF60 promotes cell cycle and lung cancer progression by regulating alternative splicing of CDC25C. Cell Rep. 42, 113041 (2023)
J.C. Marine, S.J. Dawson, M.A. Dawson, Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756 (2020)
G. Morad, B.A. Helmink, P. Sharma, J.A. Wargo, Hallmarks of response, resistance, and toxicity to immune checkpoint blockade. Cell 184, 5309–5337 (2021)
Y. Yuan, Spatial heterogeneity in the tumor microenvironment. Cold Spring Harb. Perspect. Med. 6, a026583 (2016)
L. Sun, X. Kang, C. Wang, R. Wang, G. Yang, W. Jiang, Q. Wu, Y. Wang, Y. Wu, J. Gao, et al., Single-cell and spatial dissection of precancerous lesions underlying the initiation process of oral squamous cell carcinoma. Cell Discov. 9, 28 (2023)
J. Qi, H. Sun, Y. Zhang, Z. Wang, Z. Xun, Z. Li, X. Ding, R. Bao, L. Hong, W. Jia, et al, Single-cell and spatial analysis reveal interaction of FAP(+) fibroblasts and SPP1(+) macrophages in colorectal cancer. Nat. Commun. 13, 1742 (2022)
M. Kumar, R.R. Bowers, J.R. Delaney, Single-cell analysis of copy-number alterations in serous ovarian cancer reveals substantial heterogeneity in both low- and high-grade tumors. Cell Cycle 19, 3154–3166 (2020)
Y. Luo, T. Tao, R. Tao, G. Huang, S. Wu, Single-cell transcriptome comparison of bladder cancer reveals its ecosystem. Front. Oncol. 12, 818147 (2022)
W.J. Chen, H. Cao, J.W. Cao, L. Zuo, F.J. Qu, D. Xu, H. Zhang, H.Y. Gong, J.X. Chen, J.Q. Ye, et al, Heterogeneity of tumor microenvironment is associated with clinical prognosis of non-clear cell renal cell carcinoma: a single-cell genomics study. Cell Death Dis. 13, 50 (2022)
S. Wang, G. Fan, L. Li, Y. He, N. Lou, T. Xie, L. Dai, R. Gao, M. Yang, Y. Shi, X. Han, Integrative analyses of bulk and single-cell RNA-seq identified cancer-associated fibroblasts-related signature as a prognostic factor for immunotherapy in NSCLC. Cancer Immunol. Immunother. 72, 2423–2442 (2023)
P. Ma, H.M. Amemiya, L.L. He, S.J. Gandhi, R. Nicol, R.P. Bhattacharyya, C.S. Smillie, D.T. Hung, Bacterial droplet-based single-cell RNA-seq reveals antibiotic-associated heterogeneous cellular states. Cell 186, 877–891 e814 (2023)
O.E. Rahma, F.S. Hodi, The intersection between tumor angiogenesis and immune suppression. Clin. Cancer Res. 25, 5449–5457 (2019)
S. Matsuda, A. Revandkar, T.D. Dubash, A. Ravi, B.S. Wittner, M. Lin, R. Morris, R. Burr, H. Guo, K. Seeger, et al, TGF-beta in the microenvironment induces a physiologically occurring immune-suppressive senescent state. Cell Rep. 42, 112129 (2023)
S. St Clair, J.J. Manfredi, The dual specificity phosphatase Cdc25C is a direct target for transcriptional repression by the tumor suppressor p53. Cell Cycle 5, 709–713 (2006)
K. Liu, M. Zheng, R. Lu, J. Du, Q. Zhao, Z. Li, Y. Li, S. Zhang, The role of CDC25C in cell cycle regulation and clinical cancer therapy: a systematic review. Cancer Cell Int. 20, 213 (2020)
A. Golebiewska, A.C. Hau, A. Oudin, D. Stieber, Y.A. Yabo, V. Baus, V. Barthelemy, E. Klein, S. Bougnaud, O. Keunen, et al, Patient-derived organoids and orthotopic xenografts of primary and recurrent gliomas represent relevant patient avatars for precision oncology. Acta Neuropathol. 140, 919–949 (2020)
E.R. Zanella, E. Grassi, L. Trusolino, Towards precision oncology with patient-derived xenografts. Nat. Rev. Clin. Oncol. 19, 719–732 (2022)
Acknowledgements
Thanks to the patients who provided clinical data for this medical study.
Funding
This research was supported by the Clinical Research Incubation Project of West China Hospital of Sichuan University (2018HXFH012) and the National Natural Science Foundation of China (82173251).
Author information
Authors and Affiliations
Contributions
LZ conceived the project and designed the experiments. SQM, YLW and NNC were involved in the analysis of the experiments and wrote the paper. SQM performed the bioinformatic analysis. YLW, NNC and LYZ contributed to the experiments and analyzed the data. All the authors discussed the results and reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Research Ethics Committee of West China Hospital, Sichuan University. Ethical approval for this study was provided by the Ethics Committee on Biomedical research, West China Hospital of Sichuan University on 18/12/2020.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing of interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mao, S., Wang, Y., Chao, N. et al. Integrated analysis of single-cell RNA-seq and bulk RNA-seq reveals immune suppression subtypes and establishes a novel signature for determining the prognosis in lung adenocarcinoma. Cell Oncol. (2024). https://doi.org/10.1007/s13402-024-00948-4
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-024-00948-4