Abstract
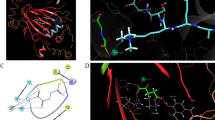
DNA methyl transferases (DNMTs) are one of the crucial epigenetic modulators associated with a wide variety of cancer conditions. Among the DNMT isoforms, DNMT1 is correlated with bladder, pancreatic, and breast cancer, as well as acute myeloid leukemia and esophagus squamous cell carcinoma. Therefore, the inhibition of DNMT1 could be an attractive target for combating cancers and other metabolic disorders. The disadvantages of the existing nucleoside and non-nucleoside DNMT1 inhibitors are the main motive for the discovery of novel promising inhibitors. Here, pharmacophore modeling, 3D-QSAR, and e-pharmacophore modeling of DNMT1 inhibitors were performed for the large fragment database screening. The resulting fragments with high dock scores were combined into molecules. The current study revealed several constitutional pharmacophoric features that can be essential for selective DNMT1 inhibition. The fragment docking and virtual screening identified 10 final hit molecules that exhibited good binding affinities in terms of docking score, binding free energies, and acceptable ADME properties. Also, the modified lead molecules (GL1b and GL2b) designed in this study showed effective binding with DNMT1 confirmed by their docking scores, binding free energies, 3D-QSAR predicted activities and acceptable drug-like properties. The MD simulation studies also suggested that leads (GL1b and GL2b) formed stable complexes with DNMT1. Therefore, the findings of this study can provide effective information for the development/identification of novel DNMT1 inhibitors as effective anticancer agents.








Similar content being viewed by others
References
Egger G, Liang G, Aparicio A, Jones PA (2004) Epigenetics in human disease and prospects for epigenetic therapy. Nature 429(6990):457–463. https://doi.org/10.1038/nature02625
Jones PA, Baylin SB (2002) The fundamental role of epigenetic events in cancer. Nat Rev Genet 3(6):415–428. https://doi.org/10.1038/nrg816
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacol 38(1):23–38. https://doi.org/10.1038/npp.2012.112
Jurkowska RZ, Jurkowski TP, Jeltsch A (2011) Structure and function of mammalian DNA methyltransferases. ChemBioChem 12(2):206–222. https://doi.org/10.1002/cbic.201000195
Hermann A, Goyal R, Jeltsch A (2004) The DNMT1 DNA-(Cytosine-C5)-Methyltransferase methylates DNA processively with high preference for hemimethylated target sites. J Biol Chem 279(46):48350–48359. https://doi.org/10.1074/jbc.M403427200
Jin B, Robertson KD (2013) DNA methyltransferases, DNA damage repair, and cancer, In: Karpf AR (ed) Epigenetic alterations in oncogenesis, Springer, New York, vol 754, pp 3–29. https://doi.org/10.1007/978-1-4419-9967-2_1
Subramaniam D, Thombre R, Dhar A, Anant S (2014) DNA methyltransferases: A novel target for prevention and therapy. Front Oncol 4:80. https://doi.org/10.3389/fonc.2014.00080
Jeltsch A (2006) Molecular enzymology of mammalian DNA methyltransferases. In: Doerfler W, Böhm P (eds) DNA methylation: basic mechanisms, Springer, Berlin, vol 301, pp 203–225. https://doi.org/10.1007/3-540-31390-7_7.
Zhang W, Xu J (2017) DNA methyltransferases and their roles in tumorigenesis. Biomark Res 20(5):1. https://doi.org/10.1186/s40364-017-0081-z
Phanus-Umporn C, Prachayasittikul V, Nantasenamat C, Prachayasittikul S, Prachayasittikul V (2020) QSAR-driven rational design of novel DNA methyltransferase 1 inhibitors. EXCLI J. 19:458–475
Medina-Franco JL, Yoo J (2013) Docking of a novel DNA methyltransferase inhibitor identified from high-throughput screening: insights to unveil inhibitors in chemical databases. Mol Divers 17(2):337–344. https://doi.org/10.1007/s11030-013-9428-z
Robertson KD (2001) DNA methylation, methyltransferases, and cancer. Oncogene 20(24):3139–3155. https://doi.org/10.1038/sj.onc.1204341
Sukur G, Uysal F, Cinar O (2023) High-fat diet induced obesity alters dnmt1 and dnmt3a levels and global DNA methylation in mouse ovary and testis. Histochem Cell Biol 159(4):339–352. https://doi.org/10.1007/s00418-022-02173-2
AmruthMaroju P, NagaMohan K (2022) DNA methyltransferases and schizophrenia: current status. In: Fukao K (ed) Psychosis - phenomenology, psychopathology and pathophysiology, IntechOpen, 2022. https://doi.org/10.5772/intechopen.98567.
Wong KK (2021) DNMT1: A key drug target in triple-negative breast cancer. Semin Cancer Biol 72:198–213. https://doi.org/10.1016/j.semcancer.2020.05.010
Dhawan D, Ramos-Vara JA, Hahn NM, Waddell J, Olbricht GR, Zheng R, Stewart JC, Knapp DW (2013) DNMT1: an emerging target in the treatment of invasive urinary bladder cancer. Semin Orig Investig 31(8):1761–1769. https://doi.org/10.1016/j.urolonc.2012.03.015
Wong KK (2020) DNMT1 as a therapeutic target in pancreatic cancer: mechanisms and clinical implications. Cell Oncol 43(5):779–792. https://doi.org/10.1007/s13402-020-00526-4
Li M, Zhang D (2022) DNA methyltransferase-1 in acute myeloid leukaemia: beyond the maintenance of DNA methylation. Ann Med 54(1):2011–2023. https://doi.org/10.1080/07853890.2022.2099578
Zhao SL, Zhu ST, Hao X, Li P, Zhang ZT (2011) Effects of DNA methyltransferase 1 inhibition on esophageal squamous cell carcinoma: the role of DNMT1 in ESCC. Diseas Esophagus 24(8):601–610. https://doi.org/10.1111/j.1442-2050.2011.01199.x
Dong E, Ruzicka WB, Grayson DR, Guidotti A (2015) DNA-Methyltransferase1 (DNMT1) binding to CpG Rich GABAergic and BDNF promoters is increased in the brain of schizophrenia and bipolar disorder patients. Schizophrenia Res 167(1–3):35–41. https://doi.org/10.1016/j.schres.2014.10.030
Chen S, Wang Y, Zhou W, Li S, Peng J, Shi Z, Hu J, Liu YC, Ding H, Lin Y, Li L, Cheng S, Liu J, Lu T, Jiang H, Liu B, Zheng M, Luo C (2014) Identifying novel selective non-nucleoside DNA methyltransferase 1 Inhibitors through docking-based virtual screening. J Med Chem 57(21):9028–9041. https://doi.org/10.1021/jm501134e
Hu C, Liu X, Zeng Y, Liu J, Wu F (2021) DNA methyltransferase inhibitors combination therapy for the treatment of solid tumor: mechanism and clinical application. Clin Epigenet 13(1):166. https://doi.org/10.1186/s13148-021-01154-x
Gnyszka A, Jastrzebski Z, Flis S (2013) DNA methyltransferase inhibitors and their emerging role in epigenetic therapy of cancer. Anticancer Res 33(20):2989–2996. https://doi.org/10.1093/jnci/dji311
Fang MZ, Chen D, Sun Y, Jin Z, Christman JK, Yang CS (2005) Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy. Clin Cancer Res 11(19 Pt 1):7033–7041. https://doi.org/10.1158/1078-0432.CCR-05-0406
Fang MZ, Wang Y, Ai N, Hou Z, Sun Y, Lu H, Welsh W, Yang CS (2003) Tea polyphenol (-)-epigallocatechin-3-gallate inhibits DNA methyltransferase and reactivates methylation-silenced genes in cancer cell lines. Cancer Res 63(22):7563–7570
Fagan RL, Cryderman DE, Kopelovich L, Wallrath LL, Brenner C (2013) Laccaic acid a is a direct, dna-competitive inhibitor of DNA methyltransferase 1. J Biol Chem 288(33):23858–23867. https://doi.org/10.1074/jbc.M113.480517
Zambrano P, Segura-Pacheco B, Perez-Cardenas E, Cetina L, Revilla-Vazquez A, Taja-Chayeb L, Chavez-Blanco A, Angeles E, Cabrera G, Sandoval K, Trejo-Becerril C, Chanona-Vilchis J, Duenas-González A (2005) A phase I study of hydralazine to demethylate and reactivate the expression of tumor suppressor genes. BMC Cancer 5:44. https://doi.org/10.1186/1471-2407-5-44
Villar-Garea A, Fraga MF, Espada J, Esteller M (2003) Procaine is a DNA-demethylating agent with growth-inhibitory effects in human cancer cells. Cancer Res 63(16):4984–4989
Cornacchia E, Golbus J, Maybaum J, Strahler J, Hanash S, Richardson B (1988) Hydralazine and procainamide inhibit T cell DNA methylation and induce autoreactivity. J Immunol 140(7):2197–2200
Siedlecki P, Boy RG, Musch T, Brueckner B, Suhai S, Lyko F, Zielenkiewicz P (2006) Discovery of two novel, small-molecule inhibitors of DNA methylation. J Med Chem 49(2):678–683. https://doi.org/10.1021/jm050844z
Brueckner B, Garcia Boy R, Siedlecki P, Musch T, Kliem HC, Zielenkiewicz P, Suhai S, Wiessler M, Lyko F (2005) Epigenetic Reactivation of tumor suppressor genes by a novel small-molecule inhibitor of human DNA methyltransferases. Cancer Res 65(14):6305–6311. https://doi.org/10.1158/0008-5472.CAN-04-2957
Datta J, Ghoshal K, Denny WA, Gamage SA, Brooke DG, Phiasivongsa P, Redkar S, Jacob ST (2009) A new class of quinoline-based DNA hypomethylating agents reactivates tumor suppressor genes by blocking DNA methyltransferase 1 activity and inducing its degradation. Cancer Res 69(10):4277–4285. https://doi.org/10.1158/0008-5472.CAN-08-3669
Gros C, Fahy J, Halby L, Dufau I, Erdmann A, Gregoire JM, Ausseil F, Vispé S, Arimondo PB (2012) DNA methylation inhibitors in cancer: recent and future approaches. Biochimie 94(11):2280–2296. https://doi.org/10.1016/j.biochi.2012.07.025
Singh N, Dueñas-González A, Lyko F, Medina-Franco JL (2009) Molecular modeling and molecular dynamics studies of hydralazine with human DNA methyltransferase 1. ChemMedChem 4(5):792–799. https://doi.org/10.1002/cmdc.200900017
Krishna S, Shukla S, Lakra AD, Meeran SM, Siddiqi MI (2017) Identification of potent inhibitors of DNA methyltransferase 1 (DNMT1) through a pharmacophore-based virtual screening approach. J Mol Graph Model 75:174–188. https://doi.org/10.1016/j.jmgm.2017.05.014
Kuck D, Singh N, Lyko F, Medina-Franco JL (2010) Novel and selective DNA methyltransferase inhibitors: docking-based virtual screening and experimental evaluation. Bioorg Med Chem 18(2):822–829. https://doi.org/10.1016/j.bmc.2009.11.050
Erlanson DA, McDowell RS, O’Brien T (2004) Fragment-based drug discovery. J Med Chem 47(14):3463–3482. https://doi.org/10.1021/jm040031v
Coronado-Posada N, Olivero-Verbel J (2019) In silico evaluation of pesticides as potential modulators of human DNA methyltransferases. SAR QSAR Environ Res 30(12):865–878. https://doi.org/10.1080/1062936X.2019.1666165
Maldonado-Rojas W, Olivero-Verbel J, Marrero-Ponce Y (2015) Computational fishing of new DNA methyltransferase inhibitors from natural products. J Mol Graph Model 60:43–54. https://doi.org/10.1016/j.jmgm.2015.04.010
Manoharan P, Ghoshal N (2018) Fragment-based virtual screening approach and molecular dynamics simulation studies for identification of BACE1 inhibitor leads. J Biomol Struct Dyn 36(7):1878–1892. https://doi.org/10.1080/07391102.2017.1337590
Murray CW, Rees DC (2009) The rise of fragment-based drug discovery. Nature Chem 1(3):187–192. https://doi.org/10.1038/nchem.217
Maestro Version 13.0, Schrodinger, LLC, New York, NY, 2021–4.
Asgatay S, Champion C, Marloie G, Drujon T, Senamaud-Beaufort C, Ceccaldi A, Erdmann A, Rajavelu A, Schambel P, Jeltsch A, Lequin O, Karoyan P, Arimondo PB, Guianvarc’h D (2014) Synthesis and evaluation of analogues of N -Phthaloyl- l -Tryptophan (RG108) as inhibitors of DNA methyltransferase 1. J Med Chem 57(2):421–434. https://doi.org/10.1021/jm401419p
Valente S, Liu Y, Schnekenburger M, Zwergel C, Cosconati S, Gros C, Tardugno M, Labella D, Florean C, Minden S, Hashimoto H, Chang Y, Zhang X, Kirsch G, Novellino E, Arimondo PB, Miele E, Ferretti E, Gulino A, Diederich M, Cheng X, Mai A (2014) Selective non-nucleoside inhibitors of human DNA methyltransferases active in cancer including in cancer stem cells. J Med Chem 57(3):701–713. https://doi.org/10.1021/jm4012627
Saavedra OM, Isakovic L, Llewellyn DB, Zhan L, Bernstein N, Claridge S, Raeppel F, Vaisburg A, Elowe N, Petschner AJ, Rahil J, Beaulieu N, MacLeod AR, Delorme D, Besterman JM, Wahhab A (2009) SAR around (l)-S-adenosyl-l-homocysteine, an inhibitor of human DNA methyltransferase (DNMT) enzymes. Bioorg Med Chem Lett 19(10):2747–2751. https://doi.org/10.1016/j.bmcl.2009.03.113
Isakovic L, Saavedra OM, Llewellyn DB, Claridge S, Zhan L, Bernstein N, Vaisburg A, Elowe N, Petschner AJ, Rahil J, Beaulieu N, Gauthier F, MacLeod AR, Delorme D, Besterman JM, Wahhab A (2009) Constrained (l-)-S-adenosyl-l-homocysteine (SAH) analogues as DNA methyltransferase inhibitors. Bioorg Med Chem Lett 19(10):2742–2746. https://doi.org/10.1016/j.bmcl.2009.03.132
Li E, Wang K, Zhang B, Guo S, Xiao S, Pan Q, Wang X, Chen W, Wu Y, Xu H, Kong X, Luo C, Chen S, Liu B (2022) Design, synthesis, and biological evaluation of novel carbazole derivatives as potent DNMT1 inhibitors with reasonable PK properties. J Enz Inhib Med Chem 37(1):1537–1555. https://doi.org/10.1080/14756366.2022.2079640
Castellano S, Kuck D, Viviano M, Yoo J, López-Vallejo F, Conti P, Tamborini L, Pinto A, Medina-Franco JL, Sbardella G (2011) Synthesis and biochemical evaluation of Δ2 -isoxazoline derivatives as dna methyltransferase 1 inhibitors. J Med Chem 54(21):7663–7677. https://doi.org/10.1021/jm2010404
Kilgore JA, Du X, Melito L, Wei S, Wang C, Chin HG, Posner B, Pradhan S, Ready JM, Williams NS (2013) Identification of DNMT1 selective antagonists using a novel scintillation proximity assay. J Biol Chem 288(27):19673–19684. https://doi.org/10.1074/jbc.M112.443895
Medina-Franco JL, López-López E, Martínez-Fernández L (2022) 7-Aminoalkoxy-quinazolines from epigenetic focused libraries are potent and selective inhibitors of DNA methyltransferase 1. Molecules 27(9):2892. https://doi.org/10.3390/molecules27092892
Yoo J, Kim JH, Robertson KD, Medina-Franco JL (2012) Molecular modeling of inhibitors of human DNA methyltransferase with a crystal structure. In: Advances in protein chemistry and structural biology; Elsevier, vol 87, pp 219–247. https://doi.org/10.1016/B978-0-12-398312-1.00008-1.
Yoo J, Medina-Franco JL (2012) Trimethylaurintricarboxylic acid inhibits human DNA methyltransferase 1: insights from enzymatic and molecular modeling studies. J Mol Model 18(4):1583–1589. https://doi.org/10.1007/s00894-011-1191-4
Medina-Franco JL, López-Vallejo F, Kuck D, Lyko F (2011) Natural products as DNA methyltransferase inhibitors: a computer-aided discovery approach. Mol Divers 15(2):293–304. https://doi.org/10.1007/s11030-010-9262-5
Gilmartin AG, Groy A, Gore ER, Atkins C, Long ER, Montoute MN, Wu Z, Halsey W, McNulty DE, Ennulat D, Rueda L, Pappalardi M, Kruger RG, McCabe MT, Raoof A, Butlin R, Stowell A, Cockerill M, Waddell I, Ogilvie D, Luengo J, Jordan A, Benowitz AB (2020) In vitro and in vivo induction of fetal hemoglobin with a reversible and selective DNMT1 inhibitor. Haematologica 106(7):1979–1987. https://doi.org/10.3324/haematol.2020.248658
Portilho NA, Deepak D, Hossain I, Sirois J, Moraes C, Pastor WA (2021) The DNMT1 inhibitor GSK-3484862 mediates global demethylation in murine embryonic stem cells. Epigenet Chromat 14(1):56. https://doi.org/10.1186/s13072-021-00429-0
Pappalardi MB, Keenan K, Cockerill M, Kellner WA, Stowell A, Sherk C, Wong K, Pathuri S, Briand J, Steidel M, Chapman P, Groy A, Wiseman AK, McHugh CF, Campobasso N, Graves AP, Fairweather E, Werner T, Raoof A, Butlin RJ, Rueda L, Horton JR, Fosbenner DT, Zhang C, Handler JL, Muliaditan M, Mebrahtu M, Jaworski JP, McNulty DE, Burt C, Eberl HC, Taylor AN, Ho T, Merrihew S, Foley SW, Rutkowska A, Li M, Romeril SP, Goldberg K, Zhang X, Kershaw CS, Bantscheff M, Jurewicz AJ, Minthorn E, Grandi P, Patel M, Benowitz AB, Mohammad HP, Gilmartin AG, Prinjha RK, Ogilvie D, Carpenter C, Heerding D, Baylin SB, Jones PA, Cheng X, King BW, Luengo JI, Jordan AM, Waddell I, Kruger RG, McCabe MT (2021) Discovery of a first-in-class reversible DNMT1-selective inhibitor with improved tolerability and efficacy in acute myeloid leukemia. Nat Cancer 2(10):1002–1017. https://doi.org/10.1038/s43018-021-00249-x
Liu Z, Xie Z, Jones W, Pavlovicz RE, Liu S, Yu J, Li P, Lin J, Fuchs JR, Marcucci G, Li C, Chan KK (2009) Curcumin is a potent dna hypomethylation agent. Bioorg Med Chem Lett 19(3):706–709. https://doi.org/10.1016/j.bmcl.2008.12.041
2D Sketcher Tool, Schrodinger, LLC, New York, NY, 2021–4.
LigPrep 13.0, Schrodinger, LLC, New York, NY, 2021–4.
Mahajan P, Suri N, Mehra R, Gupta M, Kumar A, Singh SK, Nargotra A (2017) Discovery of novel small molecule EGFR inhibitory leads by structure and ligand-based virtual screening. Med Chem Res 26:74–92. https://doi.org/10.1007/s00044-016-1728-2
Jain SV, Sonawane LV, Patil RR, Bari SB (2012) Pharmacophore modeling of some novel indole β-diketo acid and coumarin-based derivatives as HIV integrase inhibitors. Med Chem Res 21:165–173. https://doi.org/10.1007/s00044-010-9520-1
Banerjee S, Baidya SK, Adhikari N, Jha T (2022) A comparative quantitative structural assessment of benzothiazine-derived HDAC8 inhibitors by predictive ligand-based drug designing approaches. SAR QSAR Environ Res 33(12):987–1011. https://doi.org/10.1080/1062936X.2022.2155241. (Epub 2022 Dec 19 PMID: 36533308)
PHASE Version 13.0, Schrodinger, LLC, New York, NY, 2021–4.
Frimayanti N, Lee VS, Zain SM, Wahab HA, Rahman NA (2014) 2D, 3D-QSAR, and pharmacophore studies on thiazolidine-4-carboxylic acid derivatives as neuraminidase inhibitors in H3N2 influenza virus. Med Chem Res 23:1447–1453. https://doi.org/10.1007/s00044-013-0750-x
Suryanarayanan V, Sudha A, Rajamanikandan S, Vanajothi R, Srinivasan P (2013) Atom-based 3D QSAR studies on novel N-β-d-Xylosylindole derivatives as SGLT2 inhibitors. Med Chem Res 22:615–624. https://doi.org/10.1007/s00044-012-0053-7
Lanka G, Begum D, Banerjee S, Adhikari N, Yogeeswari P, Ghosh B (2023) Pharmacophore-based virtual screening 3D QSAR, Docking, ADMET, and MD simulation studies: an in silico perspective for the identification of new potential HDAC3 inhibitors. Comput Biol Med 166:107481. https://doi.org/10.1016/j.compbiomed.2023.107481
Reddy KK, Singh SK, Tripathi SK, Selvaraj C, Suryanarayanan V (2013) Shape and pharmacophore-based virtual screening to identify potential cytochrome P450 sterol 14α-Demethylase inhibitors. J Recept Signal Transduc 33(4):234–243. https://doi.org/10.3109/10799893.2013.789912
Panwar U, Singh SK (2021) In silico virtual screening of potent inhibitor to hamper the interaction between HIV-1 integrase and LEDGF/P75 interaction using e-pharmacophore modeling, molecular docking, and dynamics simulations. Comput Biol Chem 93:107509. https://doi.org/10.1016/j.compbiolchem.2021.107509
Enamine database, https://enamine.net/compound-collections/real-compounds/real-database
FCH Database, http://fchgroup.net/.
Chemdiv Database, https://www.chemdiv.scom/.
Life Chemicals Database, https://lifechemicals.com/.
Asinex Database, https://www.asinex.com/.
Protein Preparation Wizard; Epik, Schrodinger, LLC, New York, NY, 2022–1.
Receptor Grid Generation Panel 13.0, Schrodinger, LLC, New York, NY, 2022–1.
R.A. Laskowski RA, M.B. Swindells MB, (2011) LigPlot+: multiple ligand–protein interaction diagrams for drug discovery. J Chem Inf Model 51(10):2778–2786. https://doi.org/10.1021/ci200227u
Song J, Rechkoblit O, Bestor TH, Patel DJ (2011) Structure of DNMT1-DNA complex reveals a role for autoinhibition in maintenance DNA methylation. Science 331(6020):1036–1040. https://doi.org/10.1126/science.1195380
Glide Version 13.0, Schrodinger, LLC, New York, NY, 2021–4.
Elekofehinti OO, Iwaloye O, Olawale F, Chukwuemeka PO, Folorunso IM (2021) Newly designed compounds from scaffolds of known actives as inhibitors of survivin: computational analysis from the perspective of fragment-based drug design. In Silico Pharmacol 9(1):47. https://doi.org/10.1007/s40203-021-00108-8
Lanka G, Bathula R, Dasari M, Nakkala S, Bhargavi M, Somadi G, Potlapally SR (2019) Structure-based identification of potential novel inhibitors targeting FAM3B (PANDER) causing type 2 diabetes mellitus through virtual screening. J Receptor Signal Transduc 39(3):253–263. https://doi.org/10.1080/10799893.2019.1660897
Lanka G, Bhargavi M, Bathula R, Potlapally SR (2022) Targeting tribbles homolog 3 (TRIB3) protein against type 2 diabetes for the identification of potential inhibitors by in silico screening. J Ind Chem Soc 99:100531. https://doi.org/10.1016/j.jics.2022.100531
Lanka G, Bathula R, Ghosh B, Potlapally SR (2023) Identification of potential antiviral lead inhibitors against SARS-CoV-2 Main protease: structure-guided virtual screening, docking, ADME, and MD simulation based approach. Artificial Intell Chem 1:100015. https://doi.org/10.1016/j.aichem.2023.100015
Prime Version 13.0, Schrodinger, LLC, New York, NY, 2022–1.
QikProp 13.0, Schrodinger, LLC, New York, NY, 2022–1.
Desmond 13.1, Schrodinger, LLC, New York, NY, 2022–1.
Daina A, Michielin O, Zoete V (2017) SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Acknowledgements
The present work is supported by BUILDER grant from the Department of Biotechnology (DBT), India to BITS-Pilani, Hyderabad Campus (Grant No. BT/INF/22/SP42551/2021 dated 22.07.2021). Author L. Goverdhan sincerely thanks DBT-INDIA for the financial assistance as Research Associate-II under the DBT-BUILDER project. The authors are thankful to the High-Performance Computing (HPC) facility at BITS-Hyderabad Campus for providing access to run Schrodinger Software. The authors are thankful to Dr. K. Koushik, Senior Scientist, Schrodinger for his help and support during the execution of the project. The authors also acknowledge the Department of Pharmacy, BITS-Pilani, Hyderabad Campus and Department of Pharmaceutical Technology, Jadavpur University for providing research facilities to carry out the present work.
Author information
Authors and Affiliations
Contributions
GL contributed toward data curation, pharmacophore mapping, molecular docking, theoretical studies, data analysis, and manuscript drafting and writing; SB contributed toward data analysis, MD Simulation, and manuscript writing and reviewing; NA contributed toward conceptualization, data analysis, writing, editing, reviewing, overall organization of the manuscript, and supervision; BG contributed toward conceptualization, data analysis, writing, editing, reviewing, overall organization of the manuscript, and supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lanka, G., Banerjee, S., Adhikari, N. et al. Fragment-based discovery of new potential DNMT1 inhibitors integrating multiple pharmacophore modeling, 3D-QSAR, virtual screening, molecular docking, ADME, and molecular dynamics simulation approaches. Mol Divers (2024). https://doi.org/10.1007/s11030-024-10837-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-024-10837-5