Abstract
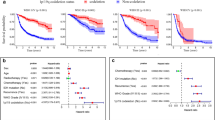
We aimed to develop and validate a predictive model for identifying long-term survivors (LTS) among glioblastoma (GB) patients, defined as those with an overall survival (OS) of more than 3 years. A total of 293 GB patients from CGGA and 169 from TCGA database were assigned to training and validation cohort, respectively. The differences in expression of immune checkpoint genes (ICGs) and immune infiltration landscape were compared between LTS and short time survivor (STS) (OS<1.5 years). The differentially expressed genes (DEGs) and weighted gene co-expression network analysis (WGCNA) were used to identify the genes differentially expressed between LTS and STS. Three different machine learning algorithms were employed to select the predictive genes from the overlapping region of DEGs and WGCNA to construct the nomogram. The comparison between LTS and STS revealed that STS exhibited an immune-resistant status, with higher expression of ICGs (P<0.05) and greater infiltration of immune suppression cells compared to LTS (P<0.05). Four genes, namely, OSMR, FMOD, CXCL14, and TIMP1, were identified and incorporated into the nomogram, which possessed good potential in predicting LTS probability among GB patients both in the training (C-index, 0.791; 0.772–0.817) and validation cohort (C-index, 0.770; 0.751–0.806). STS was found to be more likely to exhibit an immune-cold phenotype. The identified predictive genes were used to construct the nomogram with potential to identify LTS among GB patients.






Similar content being viewed by others
Data Availability
All data that support the findings of this study are openly available in TCGA (http://www.ncbi.nlm.nih.gov/geo/) and GEO database (http://www.ncbi.nlm.nih.gov/geo/).
References
Adeberg S, Bostel T, König L, Welzel T, Debus J, Combs SE (2014) A comparison of long-term survivors and short-term survivors with glioblastoma, subventricular zone involvement: a predictive factor for survival? Radiat Oncol 9:95. https://doi.org/10.1186/1748-717X-9-95. Published 2014 Apr 23
Batash R, Asna N, Schaffer P, Francis N, Schaffer M (2017) Glioblastoma multiforme, diagnosis and treatment; recent literature review. Curr Med Chem. 24(27):3002–3009. https://doi.org/10.2174/0929867324666170516123206
Burton EC, Lamborn KR, Forsyth P et al (2002) Aberrant p53, mdm2, and proliferation differ in glioblastomas from long-term compared with typical survivors. Clin Cancer Res. 8(1):180–187
Cai J, Chen Q, Cui Y et al (2018) Immune heterogeneity and clinicopathologic characterization of IGFBP2 in 2447 glioma samples. Oncoimmunology 7(5):e1426516. https://doi.org/10.1080/2162402X.2018.1426516. Published 2018 Feb 13
Cheung-Lee WL, Link AJ (2019) Genome mining for lasso peptides: past, present, and future. J Ind Microbiol Biotechnol. 46(9–10):1371–1379. https://doi.org/10.1007/s10295-019-02197-z
Das P, Puri T, Jha P et al (2011) A clinicopathological and molecular analysis of glioblastoma multiforme with long-term survival. J Clin Neurosci. 18(1):66–70. https://doi.org/10.1016/j.jocn.2010.04.050
Ding W, Zhou X, Jiang G et al (2022) Identification of prognostic biomarkers of glioblastoma based on multidatabase integration and its correlation with immune-infiltration cells. J Oncol 2022:3909030. https://doi.org/10.1155/2022/3909030. Published 2022 May 31
Fazi B, Proserpio C, Galardi S et al (2019) The expression of the chemokine CXCL14 correlates with several aggressive aspects of glioblastoma and promotes key properties of glioblastoma cells. Int J Mol Sci 20(10):2496. https://doi.org/10.3390/ijms20102496. Published 2019 May 21
Gately L, Collins A, Murphy M, Dowling A (2016) Age alone is not a predictor for survival in glioblastoma. J Neurooncol. 129(3):479–485. https://doi.org/10.1007/s11060-016-2194-x
Gately L, McLachlan SA, Philip J, Rathi V, Dowling A (2019) Molecular profile of long-term survivors of glioblastoma: a scoping review of the literature. J Clin Neurosci. 68:1–8. https://doi.org/10.1016/j.jocn.2019.08.017
Han J, Puri RK (2018) Analysis of the cancer genome atlas (TCGA) database identifies an inverse relationship between interleukin-13 receptor α1 and α2 gene expression and poor prognosis and drug resistance in subjects with glioblastoma multiforme. J Neurooncol. 136(3):463–474. https://doi.org/10.1007/s11060-017-2680-9
Hartmann C, Hentschel B, Simon M et al (2013) Long-term survival in primary glioblastoma with versus without isocitrate dehydrogenase mutations. Clin Cancer Res. 19(18):5146–5157. https://doi.org/10.1158/1078-0432.CCR-13-0017
Hertler C, Felsberg J, Gramatzki D et al (2023) Long-term survival with IDH wildtype glioblastoma: first results from the ETERNITY Brain Tumor Funders’ Collaborative Consortium (EORTC 1419). Eur J Cancer. 189:112913. https://doi.org/10.1016/j.ejca.2023.05.002
Homma T, Fukushima T, Vaccarella S et al (2006) Correlation among pathology, genotype, and patient outcomes in glioblastoma. J Neuropathol Exp Neurol. 65(9):846–854. https://doi.org/10.1097/01.jnen.0000235118.75182.94
Huang S, Cai N, Pacheco PP, Narrandes S, Wang Y, Xu W (2018) Applications of support vector machine (SVM) learning in cancer genomics. Cancer Genomics Proteomics. 15(1):41–51. https://doi.org/10.21873/cgp.20063
Jahani-Asl A, Yin H, Soleimani VD et al (2016) Control of glioblastoma tumorigenesis by feed-forward cytokine signaling. Nat Neurosci. 19(6):798–806. https://doi.org/10.1038/nn.4295
Kang K, Xie F, Wu Y et al (2021) Comprehensive exploration of tumor mutational burden and immune infiltration in diffuse glioma. Int Immunopharmacol. 96:107610. https://doi.org/10.1016/j.intimp.2021.107610
Krex D, Klink B, Hartmann C et al (2007) Long-term survival with glioblastoma multiforme. Brain. 130(Pt 10):2596–2606. https://doi.org/10.1093/brain/awm204
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD (2012) The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 28(6):882–883. https://doi.org/10.1093/bioinformatics/bts034
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P (2015) The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1(6):417–425. https://doi.org/10.1016/j.cels.2015.12.004
Louis DN, Perry A, Wesseling P et al (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 23(8):1231–1251. https://doi.org/10.1093/neuonc/noab106
Lu J, Cowperthwaite MC, Burnett MG, Shpak M (2016) Molecular predictors of long-term survival in glioblastoma multiforme patients. PLoS One 11(4):e0154313. https://doi.org/10.1371/journal.pone.0154313. Published 2016 Apr 28
Madhugiri VS, Moiyadi AV, Shetty P et al (2021) Analysis of factors associated with long-term survival in patients with glioblastoma. World Neurosurg. 149:e758–e765. https://doi.org/10.1016/j.wneu.2021.01.103
Michaelsen SR, Urup T, Olsen LR, Broholm H, Lassen U, Poulsen HS (2018) Molecular profiling of short-term and long-term surviving patients identifies CD34 mRNA level as prognostic for glioblastoma survival. J Neurooncol. 137(3):533–542. https://doi.org/10.1007/s11060-017-2739-7
Mondal B, Patil V, Shwetha SD et al (2017) Integrative functional genomic analysis identifies epigenetically regulated fibromodulin as an essential gene for glioma cell migration. Oncogene. 36(1):71–83. https://doi.org/10.1038/onc.2016.176
Moreno DA, da Silva LS, Gomes I et al (2022) Cancer immune profiling unveils biomarkers, immunological pathways, and cell type score associated with glioblastoma patients’ survival. Ther Adv Med Oncol 14:17588359221127678. https://doi.org/10.1177/17588359221127678. Published 2022 Dec 21
Noronha C, Ribeiro AS, Taipa R et al (2022) PD-L1 tumor expression is associated with poor prognosis and systemic immunosuppression in glioblastoma. J Neurooncol. 156(3):453–464. https://doi.org/10.1007/s11060-021-03907-3
Ohno M, Kitano S, Satomi K et al (2022) Assessment of radiographic and prognostic characteristics of programmed death-ligand 1 expression in high-grade gliomas. J Neurooncol. 160(2):463–472. https://doi.org/10.1007/s11060-022-04165-7
Ostrom QT, Cioffi G, Waite K, Kruchko C, Barnholtz-Sloan JS (2021) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2014–2018. Neuro Oncol 23(12 Suppl 2):iii1–iii105. https://doi.org/10.1093/neuonc/noab200
Salvati M, Cervoni L, Artico M, Caruso R, Gagliardi FM (1998) Long-term survival in patients with supratentorial glioblastoma. J Neurooncol. 36(1):61–64. https://doi.org/10.1023/a:1017926603341
Schmidt MC, Antweiler S, Urban N et al (2002) Impact of genotype and morphology on the prognosis of glioblastoma. J Neuropathol Exp Neurol. 61(4):321–328. https://doi.org/10.1093/jnen/61.4.321
Sharanek A, Burban A, Laaper M et al (2020) OSMR controls glioma stem cell respiration and confers resistance of glioblastoma to ionizing radiation. Nat Commun 11(1):4116. https://doi.org/10.1038/s41467-020-17885-z. Published 2020 Aug 17
Shinojima N, Kochi M, Hamada J et al (2004) The influence of sex and the presence of giant cells on postoperative long-term survival in adult patients with supratentorial glioblastoma multiforme. J Neurosurg. 101(2):219–226. https://doi.org/10.3171/jns.2004.101.2.0219
Shu C, Wang Q, Yan X, Wang J (2018) The TERT promoter mutation status and MGMT promoter methylation status, combined with dichotomized MRI-derived and clinical features, predict adult primary glioblastoma survival. Cancer Med. 7(8):3704–3712. https://doi.org/10.1002/cam4.1666
Sonoda Y, Kumabe T, Watanabe M et al (2009) Long-term survivors of glioblastoma: clinical features and molecular analysis. Acta Neurochir (Wien). 151(11):1349–1358. https://doi.org/10.1007/s00701-009-0387-1
Stupp R, Lukas RV, Hegi ME (2019) Improving survival in molecularly selected glioblastoma. Lancet. 393(10172):615–617. https://doi.org/10.1016/S0140-6736(18)33211-2
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Takashima Y, Kawaguchi A, Yamanaka R (2019) Promising prognosis marker candidates on the status of epithelial-mesenchymal transition and glioma stem cells in glioblastoma. Cells 8(11):1312. https://doi.org/10.3390/cells8111312. Published 2019 Oct 24
Takashima Y, Kawaguchi A, Hayano A, Yamanaka R (2019) CD276 and the gene signature composed of GATA3 and LGALS3 enable prognosis prediction of glioblastoma multiforme. PLoS One 14(5):e0216825. https://doi.org/10.1371/journal.pone.0216825. Published 2019 May 10
Tykocki T, Eltayeb M (2018) Ten-year survival in glioblastoma. A systematic review. J Clin Neurosci. 54:7–13. https://doi.org/10.1016/j.jocn.2018.05.002
Wang H, Yang F, Luo Z (2016) An experimental study of the intrinsic stability of random forest variable importance measures. BMC Bioinformatics 17:60. https://doi.org/10.1186/s12859-016-0900-5. Published 2016 Feb 3
Wang J, Liu J, Sun G et al (2019) Glioblastoma extracellular vesicles induce the tumour-promoting transformation of neural stem cells [published correction appears in Cancer Lett. 2021 Feb 1;498:245-246]. Cancer Lett 466:1–12. https://doi.org/10.1016/j.canlet.2019.09.004
Wei ST, Chiang JY, Wang HL et al (2023) Hypoxia-induced CXC chemokine ligand 14 expression drives protumorigenic effects through activation of insulin-like growth factor-1 receptor signaling in glioblastoma. Cancer Sci. 114(1):174–186. https://doi.org/10.1111/cas.15587
Weller M, van den Bent M, Preusser M et al (2021) EANO guidelines on the diagnosis and treatment of diffuse gliomas of adulthood [published correction appears in Nat Rev Clin Oncol. 2022 May;19(5):357-358]. Nat Rev Clin Oncol. 18(3):170–186. https://doi.org/10.1038/s41571-020-00447-z
Wen PY, Weller M, Lee EQ et al (2020) Glioblastoma in adults: a Society for Neuro-Oncology (SNO) and European Society of Neuro-Oncology (EANO) consensus review on current management and future directions. Neuro Oncol. 22(8):1073–1113. https://doi.org/10.1093/neuonc/noaa106
Zhang YB, Zheng SF, Ma LJ et al (2022) Elevated hexose-6-phosphate dehydrogenase regulated by OSMR-AS1/hsa-miR-516b-5p axis correlates with poor prognosis and dendritic cells infiltration of glioblastoma. Brain Sci 12(8):1012. https://doi.org/10.3390/brainsci12081012. Published 2022 Jul 30
Zhang X, Feng H, Li Z et al (2018) Application of weighted gene co-expression network analysis to identify key modules and hub genes in oral squamous cell carcinoma tumorigenesis. Onco Targets Ther 11:6001–6021. https://doi.org/10.2147/OTT.S171791. Published 2018 Sep 19
Acknowledgements
Sincere gratitude would be sent to the academic workers who were responsible for the creation, update, and maintenance of the TCGA and CGGA databases.
Funding
This work was supported by National Key R&D Program of China, Ministry of Science and Technology of the People’s Republic of China (Grant Nos. 2022YFC2407100, 2022YFC2407101).
Author information
Authors and Affiliations
Contributions
Conception/design: Xi-Lin Yang, Zheng Zeng, Fu-Quan Zhang, Xin Lian. Provision of study material or patients: Xi-Lin Yang, Chen Wang, Zheng Zeng. Collection and/or assembly of data: Xi-Lin Yang, Zheng Zeng, Guang-Yu Wang, Yun-Long Sheng. Data analysis and interpretation: Xi-Lin Yang, Zheng Zeng, Chen Wang, Yun-Long Sheng. Manuscript writing: Xi-Lin Yang. Manuscript revision: Fu-Quan Zhang, Xin Lian. Final approval of manuscript: all authors.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, XL., Zeng, Z., Wang, C. et al. Predictive Model to Identify the Long Time Survivor in Patients with Glioblastoma: A Cohort Study Integrating Machine Learning Algorithms. J Mol Neurosci 74, 48 (2024). https://doi.org/10.1007/s12031-024-02218-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12031-024-02218-2