Abstract
Background and aims
The global agricultural importance of sugarcane highlights the crucial need to understand the dynamics of rhizosphere microbiota for maximizing yields. Guangxi, a crucial sugarcane production region in China, boasts diverse landscapes. This study aims to identify microbial variations in wild soil across different terrains and to elucidate differences between planted and wild soils within plains sugarcane rhizospheres.
Methods
A comprehensive examination of soil microbiota was conducted in wild sugarcane rhizospheres spanning mountains, hills, and plain, alongside planted sugarcane rhizospheres in the plains of Guangxi, China. A total of 75 soil samples underwent high-throughput sequencing. The analysis employed microbial diversity assessment, species correlation, and functional profiling.
Results
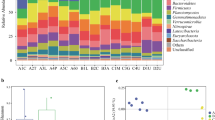
The analysis unveiled distinct disparities in microbiota composition among various wild soil topographies and between planting and wild soils within plains sugarcane rhizospheres. Notably, certain microorganisms, such as Nitrospira and Streptomyces in hills, Mesorhizobium thiogangeticum in mountain, and Clonostachys rosea and Stachybotrys bisbyi in plain, exhibited significantly higher proportions. Planting soil demonstrated lower richness and diversity of bacteria compared to wild soil. Exclusive identification of microbial species such as Lysobacter niastensis, Vicinamibacter silvestris, and Aspergillus aculeatus in wild soils suggests potential benefits for sugarcane growth. Functional profiling indicated enriched metabolic pathways in wild soils, particularly those associated with nutrient cycling and organic matter degradation.
Conclusion
This study offers a comparative analysis of sugarcane rhizosphere microbiota, highlighting potentially beneficial microbial species. These insights advance agricultural microbiology and soil science, providing a foundation for sustainable improvements in sugarcane agriculture and crop productivity.






Similar content being viewed by others
Data availability
The sequence data have been submitted to the GenBank Sequence Read Archive (accession number PRJNA1020593).
References
Bharti R, Grimm DG (2021) Current challenges and best-practice protocols for microbiome analysis. Brief Bioinform 22(1):178–193
Bolyen E et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857
Chamkhi I, El Omari N, Balahbib A, El Menyiy N, Benali T, Ghoulam C (2022) Is the rhizosphere a source of applicable multi-beneficial microorganisms for plant enhancement? Saudi J Biol Sci 29(2):1246–1259
Chen S, Wang Y, Gao J, Chen X, Qi J, Peng Z, Chen B, Pan H, Liang C, Liu J, Wang Y, Wei G, Jiao S (2023) Agricultural tillage practice and rhizosphere selection interactively drive the improvement of soybean plant biomass. Plant Cell Environ 46(11):3542–3557
Coleman-Derr D, Desgarennes D, Fonseca-Garcia C, Gross S, Clingenpeel S, Woyke T, North G, Visel A, Partida-Martinez LP, Tringe SG (2016) Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. New Phytol 209(2):798–811
Deng Q, Yu T, Zeng Z, Ashraf U, Shi Q, Huang S, Lian T, Chen J, Muzaffar W, Shen W (2021) Silicon application modulates the growth, rhizosphere soil characteristics, and bacterial community structure in sugarcane. Front Plant Sci 12:710139
Dillon RJ, Buckling CTVA, Charnley AK (2005) Diversity of locust gut bacteria protects against pathogen invasion. Ecol Lett 8(12):1291–1298
Dombrowski N, Seitz KW, Teske AP, Baker BJ (2017) Genomic insights into potential interdependencies in microbial hydrocarbon and nutrient cycling in hydrothermal sediments. Microbiome 5(1):106
Douglas GM, Maffei VJ, Zaneveld JR, Yurgel SN, Brown JR, Taylor CM, Huttenhower C, Langille MGI (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38(6):685–688
Emmett BD, Buckley DH, Drinkwater LE (2020) Plant growth rate and nitrogen uptake shape rhizosphere bacterial community composition and activity in an agricultural field. New Phytol 225(2):960–973
Fang X, Wang H, Zhao L, Wang M, Sun M (2022) Diversity and structure of the rhizosphere microbial communities of wild and cultivated ginseng. BMC Microbiol 22(1):2
Huber KJ, Geppert AM, Wanner G, Fösel BU, Wüst PK, Overmann J (2016) The first representative of the globally widespread subdivision 6 Acidobacteria,Vicinamibacter silvestris gen. Nov., sp. nov., isolated from subtropical savannah soil. Int J Syst Evol Microbiol 66(8):2971–2979
Ishida JK, Bini AP, Creste S, Van Sluys MA (2022) Towards defining the core Saccharum microbiome: input from five genotypes. BMC Microbiol 22(1):193
Khan A, Jiang H, Bu J, Adnan M, Gillani SW, Zhang M (2022) An insight to rhizosphere bacterial community composition and structure of consecutive winter-initiated sugarcane ratoon crop in southern China. BMC Plant Biol 22(1):74
Kim H, Park Y-H, Yang JE, Kim H-S, Kim S-C, Oh E-J, Moon J, Cho W, Shin W, Yu C (2022) Analysis of major bacteria and diversity of surface soil to discover biomarkers related to soil health. Toxics 10(3):117
Kirk JL, Beaudette LA, Hart M, Moutoglis P, Klironomos JN, Lee H, Trevors JT (2004) Methods of studying soil microbial diversity. J Microbiol Methods 58(2):169–188
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41(1):e1
Koistinen J, Sjöblom M, Spilling K (2020) Total nitrogen determination by a spectrophotometric method. Methods Mol Biol 1980:81–86
Li X, Han S, Wang G, Liu X, Amombo E, Xie Y, Fu J (2017) The fungus aspergillus aculeatus enhances salt-stress tolerance, metabolite accumulation, and improves forage quality in perennial ryegrass. Front Microbiol 8:1664
Li G, Liu P, Zhao J, Su L, Zhao M, Jiang Z, Zhao Y, Yang X (2023a) Correlation of microbiomes in “plant-insect-soil” ecosystem. Front Microbiol 14:1088532
Li X, Zhang T, Xue Y, Xu X, Cui X, Fu J (2023b) Aspergillus aculeatus enhances nutrient uptake and forage quality in bermudagrass by increasing phosphorus and potassium availability. Front Plant Sci 14:1165567
Liu Q, Pang Z, Liu Y, Fallah N, Hu C, Lin W, Yuan Z (2023) Rhizosphere fungal dynamics in sugarcane during different growth stages. Int J Mol Sci 24(6):5701
Marques JM, da Silva TF, Vollu RE, Blank AF, Ding GC, Seldin L, Smalla K (2014) Plant age and genotype affect the bacterial community composition in the tuber rhizosphere of field-grown sweet potato plants. FEMS Microbiol Ecol 88(2):424–435
Meena MR, Appunu C, Arun Kumar R, Manimekalai R, Vasantha S, Krishnappa G, Kumar R, Pandey SK, Hemaprabha G (2022) Recent advances in sugarcane genomics, physiology, and phenomics for superior agronomic traits. Front Genet 13:854936
Moneda APC, de Carvalho LAL, Teheran-Sierra LG, Funnicelli MIG, Pinheiro DG (2022) Sugarcane cultivation practices modulate rhizosphere microbial community composition and structure. Sci Rep 12(1):19174
Mori K, Kamagata Y (2014) The challenges of studying the anaerobic microbial world. Microbes Environ 29(4):335–337
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30(21):3123–3124
Rudrappa T, Czymmek KJ, Paré PW, Bais HP (2008) Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiol 148(3):1547–1556
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Seccareccia I, Kost C, Nett M (2015) Quantitative analysis of Lysobacter predation. Appl Environ Microbiol 81(20):7098–7105
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60
Turrini P, Artuso I, Tescari M, Lugli GA, Frangipani E, Ventura M, Visca P (2021) Draft genome sequence and secondary metabolite biosynthetic potential of the Lysobacter niastensis type strain DSM 18481. Microbiol Resour Announc 10(1):e01296–e01220
van Elsas JD, Chiurazzi M, Mallon CA, Elhottova D, Kristufek V, Salles JF (2012) Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc Natl Acad Sci USA 109(4):1159–1164
Walkley AJ, Black IA (1934) An examination of the degtjareff method for determining soil organic matter, and a proposed modification of the chromic acid titration method. Soil Sci 37:29–38
Wang S, Tian H, Liu J, Pan S (2003) Pattern and change of soil organic carbon storage in China: 1960s-1980s. Tellus Ser B Chem Phys Meteorol 55(2):416–427
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73(16):5261–5267
Xie Y, Li X, Huang X, Han S, Amombo E, Wassie M, Chen L, Fu J (2019) Characterization of the cd-resistant fungus aspergillus aculeatus and its potential for increasing the antioxidant activity and photosynthetic efficiency of rice. Ecotoxicol Environ Saf 171:373–381
Yuan J, Wen T, Zhang H, Zhao M, Penton CR, Thomashow LS, Shen Q (2020) Predicting disease occurrence with high accuracy based on soil macroecological patterns of fusarium wilt. ISME J 14(12):2936–2950
Yuan Z, Liu Q, Pang Z, Fallah N, Liu Y, Hu C, Lin W (2022) Sugarcane rhizosphere bacteria community migration correlates with growth stages and soil nutrient. Int J Mol Sci 23(18):10303
Zhang Y, Shen H, He X, Thomas BW, Lupwayi NZ, Hao X, Thomas MC, Shi X (2017) Fertilization shapes bacterial community structure by alteration of soil pH. Front Microbiol 8:1325
Zou R, Wang Y, Duan M, Guo M, Zhang Q, Zheng H (2021) Dysbiosis of gut fungal microbiota in children with autism spectrum disorders. J Autism Dev Disord 51(1):267–275
Zou YW, Zhang JH, Ning DH, Zeng YL, Li WQ, Xia CY, Zhang H, Deng HP (2022) Comparative analyses of rhizosphere bacteria along an elevational gradient of Thuja sutchuenensis. Front Microbiol 13:881921
Acknowledgements
This work was supported by Guangxi Key Laboratory of Sugarcane Genetic Improvement (21-238-16-K-03-01) to B.W., National Natural Science Foundation of China (32160486) to X.L., Research Start-up Funding of Guangxi Academy of Sciences (2021YBJ704) to Y.Q.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Responsible Editor: Eric Paterson.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, B., Liu, X., Qi, Y. et al. Sugarcane rhizosphere microbiota: exploring diversity across varied topographies and growth environments. Plant Soil (2024). https://doi.org/10.1007/s11104-024-06688-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11104-024-06688-6